sklearn_extra.cluster.CommonNNClustering¶
- class sklearn_extra.cluster.CommonNNClustering(eps=0.5, *, min_samples=5, metric='euclidean', metric_params=None, algorithm='auto', leaf_size=30, p=None, n_jobs=None)[source]¶
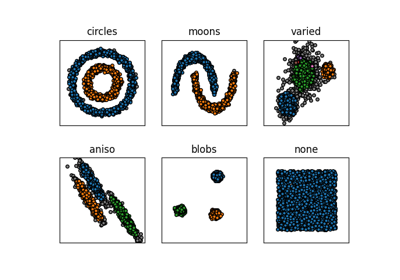
Density-Based common-nearest-neighbors clustering.
Read more in the User Guide.
- Parameters:
- epsfloat, default=0.5
The maximum distance between two samples for one to be considered as in the neighborhood of the other. This is not a maximum bound on the distances of points within a cluster. The clustering will use min_samples within eps as the density criterion. The lower eps, the higher the required sample density.
- min_samplesint, default=5
The number of samples that need to be shared as neighbors for two points being part of the same cluster. The clustering will use min_samples within eps as the density criterion. The larger min_samples, the higher the required sample density.
- metricstring, or callable, default=’euclidean’
The metric to use when calculating distance between instances in a feature array. If metric is a string or callable, it must be one of the options allowed by
sklearn.metrics.pairwise_distances()for its metric parameter. If metric is “precomputed”, X is assumed to be a distance matrix and must be square. X may be a Glossary, in which case only “nonzero” elements may be considered neighbors.- metric_paramsdict, default=None
Additional keyword arguments for the metric function.
- algorithm{‘auto’, ‘ball_tree’, ‘kd_tree’, ‘brute’},
default=’auto’ The algorithm to be used by
NearestNeighborsto compute pointwise distances and find nearest neighbors.- leaf_sizeint, default=30
Leaf size passed to tree
NearestNeighborsdepending on algorithm. This can affect the speed of the construction and query, as well as the memory required to store the tree. The optimal value depends on the nature of the problem.- pfloat, default=None
The power of the Minkowski metric to be used to calculate distance between points.
- n_jobsint, default=None
The number of parallel jobs to run. None means 1 unless in a
joblib.parallel_backendcontext. -1 means using all processors. See Glossary for more details.
See also
commonnnA function interface for this cluster algorithm.
sklearn.cluster.DBSCANA similar clustering providing a different notion of the point density. The implementation is (like this present
CommonNNClusteringimplementation) optimized for speed.sklearn.cluster.OPTICSA similar clustering at multiple values of eps. The implementation is optimized for memory usage.
Notes
This implementation bulk-computes all neighborhood queries, which increases the memory complexity to \(O(n ⋅ n_n)\) where \(n_n\) is the average number of neighbors, similar to the present implementation of
sklearn.cluster.DBSCAN. It may attract a higher memory complexity when querying these nearest neighborhoods, depending on the algorithm.One way to avoid the query complexity is to pre-compute sparse neighborhoods in chunks using
NearestNeighbors.radius_neighbors_graphwith mode=’distance’, then using metric=’precomputed’ here.sklearn.cluster.OPTICSprovides a similar clustering with lower memory usage.References
B. Keller, X. Daura, W. F. van Gunsteren “Comparing Geometric and Kinetic Cluster Algorithms for Molecular Simulation Data” J. Chem. Phys., 2010, 132, 074110.
O. Lemke, B.G. Keller “Density-based Cluster Algorithms for the Identification of Core Sets” J. Chem. Phys., 2016, 145, 164104.
O. Lemke, B.G. Keller “Common nearest neighbor clustering - a benchmark” Algorithms, 2018, 11, 19.
Examples
>>> from sklearn_extra.cluster import CommonNNClustering >>> import numpy as np >>> X = np.array([[1, 2], [2, 2], [2, 3], [8, 7], [8, 8], [25, 80]]) >>> clustering = CommonNNClustering(eps=3, min_samples=0).fit(X) >>> clustering.labels_ array([ 0, 0, 0, 1, 1, -1])
- Attributes:
- labels_ndarray of shape (n_samples)
Cluster labels for each point in the dataset given to fit(). Noisy samples are given the label -1.