sklearn_extra.cluster.CLARA¶
- class sklearn_extra.cluster.CLARA(n_clusters=8, metric='euclidean', init='build', max_iter=300, n_sampling=None, n_sampling_iter=5, random_state=None)[source]¶
CLARA clustering.
Read more in the User Guide. CLARA (Clustering for Large Applications) extends k-medoids approach for a large number of objects. This algorithm use a sampling approach.
- Parameters:
- n_clustersint, optional, default: 8
The number of clusters to form as well as the number of medoids to generate.
- metricstring, or callable, optional, default: ‘euclidean’
What distance metric to use. See :func:metrics.pairwise_distances
- max_iterint, optional, default300
Specify the maximum number of iterations when fitting PAM. It can be zero in which case only the initialization is computed.
- n_samplingint or None, optional, defaultNone
Size of the sampled dataset at each iteration. sampling-size a trade-off between complexity and efficiency. If None, then sampling-size is set to min(sample_size, 40 + 2 * self.n_clusters) as suggested by the authors of the algorithm. must be smaller than sample_size.
- n_sampling_iterint, optional, default5
Number of different samples that have to be done, or number of iterations.
- random_stateint, RandomState instance or None, optional
Specify random state for the random number generator. Used to initialise medoids when init=’random’.
See also
KMedoidsCLARA is a variant of KMedoids that use sub-sampling scheme as such if the dataset is sufficiently small, KMedoids is preferable.
Notes
Contrary to KMedoids, CLARA is linear in N the sample size for both the spacial and time complexity. On the other hand, it scales quadratically with n_sampling.
Examples
>>> from sklearn_extra.cluster import CLARA >>> import numpy as np >>> from sklearn.datasets import make_blobs >>> X, _ = make_blobs(centers=[[0,0],[1,1]], n_features=2,random_state=0) >>> clara = CLARA(n_clusters=2, random_state=0).fit(X) >>> clara.predict([[0,0], [4,4]]) array([0, 1]) >>> clara.inertia_ 122.44919397611667
- Attributes:
- cluster_centers_array, shape = (n_clusters, n_features)
or None if metric == ‘precomputed’
Cluster centers, i.e. medoids (elements from the original dataset)
- medoid_indices_array, shape = (n_clusters,)
The indices of the medoid rows in X
- labels_array, shape = (n_samples,)
Labels of each point
- inertia_float
Sum of distances of samples to their closest cluster center.
Examples using sklearn_extra.cluster.CLARA¶
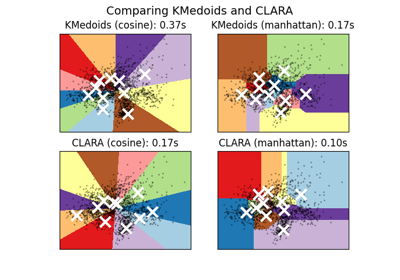
A demo of K-Medoids vs CLARA clustering on the handwritten digits data